Atlas solution¶
The Atlas Solutions module evaluates adaptation strategies designed to mitigate the impacts of climate change on crop suitability.
One example of such a strategy is the simulation of improved crop varieties, specifically cultivars bre for enhanced tolerance to drought or heat stress. This module quantifies how adopting these resilient varieties alters the suitability landscape compared to baseline crops.
1. Membership functions¶
CropSuite uses a fuzzy-logic approach based on Liebig’s Law of the Minimum. Suitability is assessed using crop-specific that map abiotic conditions (e.g., temperature, precipitation, soil properties) to a suitability score ranging from 0 (Not Suitable) to 1 (Highly Suitable).
Most baseline membership functions are defined according to Sys et al. (1993).
Here are some examples of the original membership functions:
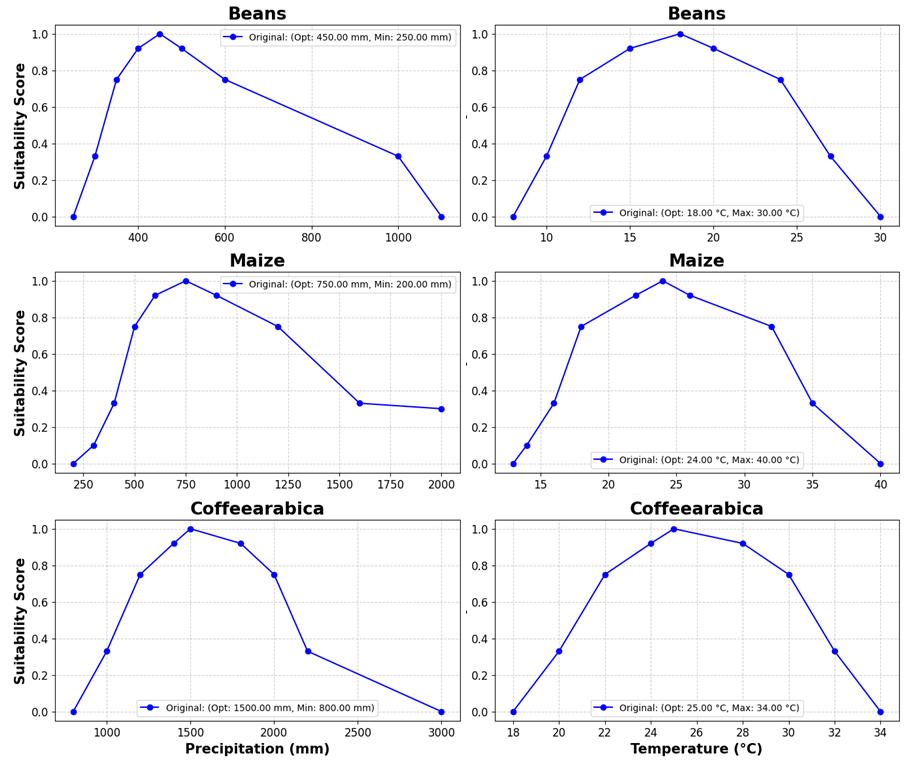
Figure 1. Membership functions
Each plot can be obtained using the following code
from src.membership_functions import CropSensitivity
cs = CropSensitivity('coffeearabica', 'plant_params/available')
cs.plot_solutions_profiles(variable='prec', labelsize = 15, add_solution= False, figsize = (8, 4))
Simulating Improved Varieties¶
To represent the effect of improved cultivars, this module allows to modify the membership functions for temperature and precipitation.
-
Heat Tolerance: The maximum temperature threshold of the function is increased (multiplied by a percentage factor), assuming the new variety can tolerate higher temperatures.
-
Drought Tolerance: For drought-tolerant varieties, the adjustment is done on the minimun suitability values, increasing the tolerance of the plant to water scarcity.
2. Visualization Tools¶
The module allows you to visualize how these "Solutions" modify the crop requirements.
Plotting Baseline vs. Adapted Curves¶
The following code demonstrates how to modify and plot the membership functions.
from solutions.membership_functions import CropSensitivity
cs = CropSensitivity('beans', 'plant_params/available')
cs.read_crop_configuration()
solution_parameter = {}
# set meteorological variable
variable = 'prec'
# modify the membership functions using a precentage
if variable == 'temp':
parameter_suit, new_param_vals = cs.create_new_max_parameter_values( parameter=variable,new_max=11.7, percentage = True)
else:
parameter_suit, new_param_vals = cs.multiply_suit_vals(parameter='prec',perc_factor=50)
solution_parameter.update({f'{variable}_1':new_param_vals})
# plot
cs.plot_solutions_profiles(scenario_values=solution_parameter, suit_values = parameter_suit, labelsize = 15, add_solution= True, figsize = (8, 4))
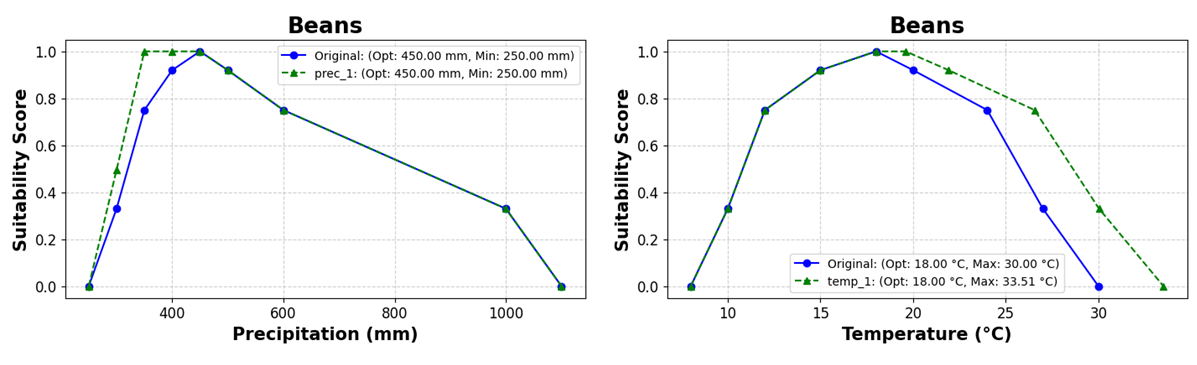
Figure 1. Membership functions modified for droguth (left) and heat (rigth) tolerant varieties
3. Automated Workflow¶
To obtain the crop suitability maps can be obtained automatically by changing the configuration files
Define Response Factors¶
The yaml file yaml_configurations\response_functions.yaml contains the percentage adjustment factors used to modify the membership functions for each crop–solution combination. These values are based on scientific evidence regarding stress-tolerant varieties (see the Evidence section). These values are based on scientific evidence regarding stress-tolerant varieties (see the Evidence section).
It also defines the codes used to identify crops and solution types. Currently there are available 26 crops.
This is an example of the file's content
SOLUTIONS_TYPE:
ST1: 'Croop Breeding'
SOLUTIONS_CODE:
s1: 'Drought-tolerant varieties'
s2: 'Heat-tolerant varieties'
CROPS:
c1: maize
c2: millet
c3: wheat
c4: rice
ST1:
s1:
c1: [15]
c2: [30]
c3: [50]
c4: [24]
s2:
c1: [9.1]
c2: [23.8]
c3: [11.8]
c4: [7]
c5: [11.7]
Modify the configuration file¶
To execute a batch run, modify yaml_configurations/general_config_solutions.yaml. Update the SOLUTIONS section to specify which strategies and crops to process.
Three elements can be configured:
-
solutions_path → path to the YAML file containing modification factors
-
solution_codes → list of solution types to apply
-
crop_codes → list of crops to evaluate
SOLUTIONS:
solutions_path: 'yaml_configurations/response_functions.yaml'
solution_codes: ['s1', 's2']
crop_codes: ['c1', 'c2', 'c3' ]
Execution¶
Once the YAML is configured, run the full pipeline from the command line: